A research team led by Professor Xinghua Pan and Chair Professor Paul Kwong Hang Tam from the Precision Regenerative Medicine Research Center (PRMRC), Medical Sciences Division (MSD), Macau University of Science and Technology (MUST), in collaboration with the researchers from the School of Basic Medical Sciences, Zhujiang Hospital and Nanfang Hospital of Southern Medical University, has recently achieved a significant breakthrough in the field of hepatocellular carcinoma (HCC). Firstly, based on single-nucleus RNA sequencing (snRNA-seq) of small HCC samples, they identified 10 hepatocyte subpopulations and their different underlying molecular profile and risk for HCC progression. These founding were validated through bulk RNA sequencing (bRNA-seq) of relatively larger sample cohort of their own. Secondly, they designed a novel quantile-based scoring method, which effectively consolidated large-scale transcriptomic data in varied forms from multiple independent cohorts. Then leveraging machine learning techniques and the new message obtained above, they established a disease diagnosis and treatment model. Thirdly, they proposed a new HCC classification system based on molecular regulatory mechanisms, and they predicted potential therapeutic drugs tailored to different HCC subclass. This groundbreaking research has unveiled new mechanisms, therapeutic targets, and classification for hepatocellular carcinoma, providing fresh theoretical foundations for the early diagnosis and treatment of HCC. The study, entitle "Deciphering the Oncogenic Landscape of Hepatocytes Through Integrated Single-Nucleus and Bulk RNA-Seq of Hepatocellular Carcinoma", was published in Advanced Science (Impact Factor 14.3, JCR Q1).
Unveils new insights of Hepatocellular Carcinoma, addressing a major treatment challenge
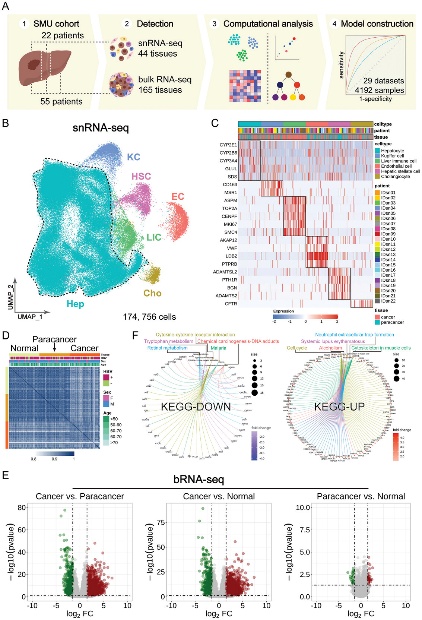 HCC is the sixth most common cancer worldwide and the third leading cause of cancer-related deaths. Its asymptomatic nature in the early stages, limited treatment options in advanced stages, high recurrence rates, and poor prognosis pose significant challenges for clinical diagnosis and treatment. In recent years, advancements in genomics have brought new hope for HCC prevention and treatment. Along with these, bRNA-seq provides an overall transcriptomic profile of cancer tissues but often masks the genetic characteristics of different cellular subpopulations. In contrast, single cell transcriptome sequencing, conventionally known as scRNA-seq, offers high-resolution insights into the transcriptomic information of individual cells, accurately reflecting cellular-level functional heterogeneity. However, due to the large size and fragile nature of hepatocytes, the conventional whole-cell scRNA-seq methods tend to enrich the microenvironmental cells while depleting most of the parenchymal cells in liver, ie hepatocytes. Consequently, the understanding of the hepatocyte heterogeneity remain limited. The single-nucleus RNA sequencing (snRNA-seq) captures all cell nuclei within a sample in a unbiased way, thus more accurately reflecting the natural composition of cell types in the tissue, including the intrinsic characteristics of hepatocytes and microenvironment cells in HCC cells. Combining both snRNA-seq and bRNA-seq, the research team conducted a comprehensive analysis of carcinogenesis and molecular regulation using a cohort of 77 HCC patients, identifying the key cellular subpopulations, genes, and pathways, as well as a series of novel diagnostic and therapeutic targets. To further validate and expand their findings, the researchers developed a quantile-based scoring method that effectively integrated transcriptomic information from 4,192 samples across 29 HCC cohorts. By constructing a Quantile Distribution Model (QDM) for HCC, they verified and extended the experimental results. Building on this foundation, the study redefined HCC into three subclasses with molecular network profiles: metabolic-HCC, inflammatory-HCC, and matrix-HCC. The team elucidated the distinct clinical characteristics of each HCC subclass and identified their corresponding priority candidate therapeutic drugs. This research not only advances the understanding of HCC pathogenesis but also provides new theoretical foundations and molecular targets for early diagnosis and personalized treatment, marking a significant milestone in the field of HCC.
HCC is the sixth most common cancer worldwide and the third leading cause of cancer-related deaths. Its asymptomatic nature in the early stages, limited treatment options in advanced stages, high recurrence rates, and poor prognosis pose significant challenges for clinical diagnosis and treatment. In recent years, advancements in genomics have brought new hope for HCC prevention and treatment. Along with these, bRNA-seq provides an overall transcriptomic profile of cancer tissues but often masks the genetic characteristics of different cellular subpopulations. In contrast, single cell transcriptome sequencing, conventionally known as scRNA-seq, offers high-resolution insights into the transcriptomic information of individual cells, accurately reflecting cellular-level functional heterogeneity. However, due to the large size and fragile nature of hepatocytes, the conventional whole-cell scRNA-seq methods tend to enrich the microenvironmental cells while depleting most of the parenchymal cells in liver, ie hepatocytes. Consequently, the understanding of the hepatocyte heterogeneity remain limited. The single-nucleus RNA sequencing (snRNA-seq) captures all cell nuclei within a sample in a unbiased way, thus more accurately reflecting the natural composition of cell types in the tissue, including the intrinsic characteristics of hepatocytes and microenvironment cells in HCC cells. Combining both snRNA-seq and bRNA-seq, the research team conducted a comprehensive analysis of carcinogenesis and molecular regulation using a cohort of 77 HCC patients, identifying the key cellular subpopulations, genes, and pathways, as well as a series of novel diagnostic and therapeutic targets. To further validate and expand their findings, the researchers developed a quantile-based scoring method that effectively integrated transcriptomic information from 4,192 samples across 29 HCC cohorts. By constructing a Quantile Distribution Model (QDM) for HCC, they verified and extended the experimental results. Building on this foundation, the study redefined HCC into three subclasses with molecular network profiles: metabolic-HCC, inflammatory-HCC, and matrix-HCC. The team elucidated the distinct clinical characteristics of each HCC subclass and identified their corresponding priority candidate therapeutic drugs. This research not only advances the understanding of HCC pathogenesis but also provides new theoretical foundations and molecular targets for early diagnosis and personalized treatment, marking a significant milestone in the field of HCC.
Impact and Significance
HCC poses a major threat to human health worldwide, particularly among Asian populations. Leveraging cutting-edge single-cell experimental techniques and advanced artificial intelligence-based data analysis, this study elucidated the molecular characteristics and functional roles of different hepatocyte subtypes in HCC and their correlations with clinical indicators. Utilizing the unique features of these hepatocyte subtypes, the research team developed a high-performance diagnostic model for HCC based on multi-center datasets and identified potential disease-associated biomarkers. This study provides the most comprehensive human HCC transcriptomic resource to date and establishes a novel classification system based on hepatocyte subtypes and regulatory networks. These advancements offer a solid scientific foundation and innovative insights for further explore of the mechanisms of HCC development, devising targeted therapeutic strategies.
About the Research Team
This study was conducted by a team of researchers from the Macau University of Science and Technology (MUST) and the Southern Medical University, with support from National Natural Science Foundation of China (NSFC), Guangdong Foundation for Basic and Applied Basic Research, Faculty Research Grant of Macau University of Science and technology (FRG) and Macau Science and Technology Development Fund (FDCT). The research team from PRMRC at the MSD of MUST. Was led by Director Chair Professor Paul Kwong Hang, Assistant Director Yen Chen and Professor Xinghua Pan, with Professor Xinghua Pan serving as the chief corresponding author of this journal.
Link to the study: https://advanced.onlinelibrary.wiley.com/doi/10.1002/advs.202412944




























